Professor Itaru Hamachi, Lecturer Tomonori Tamura, and Assistant Professor Masaki Tsuchiya of Kyoto University have developed a new technology called O-ClickFC (Organelle-selective click labeling with flow cytometry), which can, at a very high speed, analyze the metabolic state of lipids in cells.
Conventional analysis of cellular lipids mainly involves radiometric analysis and mass spectrometry of cell extracts collected in large quantities. This method is time-consuming and labor-intensive and burdensome when analyzing a large number of samples. The issue needs to be addressed to elucidate the relationship between 20,000 types of human genes and lipid metabolism. In this study, the researchers developed O-ClickFC, a technology that converts the abundance and spatial distribution of lipids in living cells into simple fluorescent signal information and analyzes them at an ultra-high speed (10,000 cells per second). This is done using a unique click reaction(1) that can label lipids with fluorescent dyes in living cells. By combining this technology with “genome editing”(2), it is possible to select cells with abnormal lipid metabolism from a cell population with mutations in all human genes and identify the causative genes. As a demonstrative experiment, the research group identified 49 genes important for the metabolism of phosphatidylcholine (PC)(3), a major component of human lipids, and discovered many novel genes, including FLVCR1, in the process. From a thorough analysis, it was found that FLVCR1 played a role in the uptake of choline, a nutrient essential for normal bodily functions and human health. Furthermore, the researchers elucidated part of the pathogenesis mechanism in mutant FLVCR1, which causes hereditary neurological disease and loss of choline uptake.
It has become evident over the years that metabolic abnormalities are the source of diseases such as cancer, obesity, and diabetes. O-ClickFC may be thereby applied for the analysis of not only lipids but also various metabolites such as sugars and amino acids to identify the genetic factors that link pathogenesis and metabolic abnormalities, as well as the discovery of candidate molecules as therapeutic targets.
(1) Click reaction
Substance synthesis reaction: Under physiological conditions such as in vivo or in cells, two molecules can be easily, rapidly, and efficiently bound. As a representative example, a coupling reaction of two molecules called azide and alkyne is often used. In this technology, azide is introduced into lipids through metabolic pathways, and fluorescent labeling of the lipids is performed by allowing them to react with fluorescent dyes containing alkyne.
(2) Genome editing, CRISPR-KO screening
Genetic modification technique: Using a highly efficient DNA-cleaving enzyme system called CRISPR/Cas9, it is possible to introduce mutations into the cell genome (DNA in which all genetic information is written) and delete the function of specific genes (KO: Knockout). CRISPR-KO screening uses cell populations (libraries) that are deficient in one gene per cell, and in the case of the human genome, about 20,000 genes are targeted. By selecting a small percentage of the cells exhibiting the desired biological characteristics from a cell library of about 100 million, it is possible to identify the characteristic gene mutations.
(3) Phosphatidylcholine (PC)
PC is a major component of lipids in mammalian cells, including those of humans. It is a base molecule that forms the cell membrane and is involved in the metabolic balance of other types of lipids (such as triglycerides). Choline, which is the source of PC, is an essential nutrient that must be externally ingested through diet or other means because cells cannot produce it on their own. A choline deficiency or defective PC synthesis causes severe damage to the cells. In humans, retinal diseases, muscle diseases, lipodystrophy, and the like represent a few hereditary diseases resulting from poor phosphatidylcholine synthesis.
This study was supported by JST under the ERATO program: HAMACHI Innovative Molecular Technology for Neuroscience and PRESTO program: "A flow cytometry-based method using chemical labeling for genetic dissection of cellular lipid dynamics" in the research area of "Dynamic supra-assembly of biomolecular systems".
-
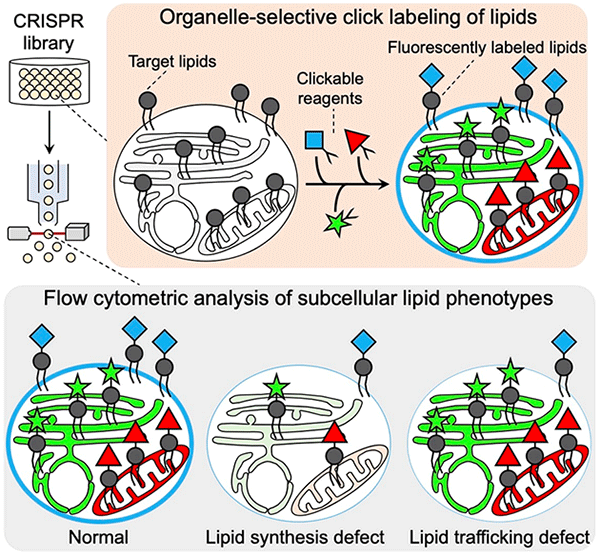
Figure1: O-ClickFC for high-throughput analysis of single-cell lipid metabolism at the organelle level O-ClickFC is applicable to CRISPR screening for discovery of genetic factors underlying human lipid metabolism. Target lipids in live cells are fluorescently labeled via spatially limited click reactions using different-colored organelle-targeting reagents. This allows for flow cytometric high-throughput analysis of their lipid contents and distribution at the organelle level on the basis of fluorescent intensity and spectrum of labeled lipids. Fluorescent patterns on labeled lipids in individual cells reflect the metabolic status such as defects in lipid biosynthesis or trafficking caused by CRISPR gene editing, thereby enabling large-scale genetic screening based on subcellular lipid phenotypes. ©JST
Program Information
- JST ERATO
- Research Area: Life Innovation
- Research theme: HAMACHI Innovative Molecular Technology for Neuroscience
- JST PRESTO
- Research Area: Dynamic supra-assembly of biomolecular systems
- Research Theme “A flow cytometry-based method using chemical labeling for genetic dissection of cellular lipid dynamics”
Journal Information
Masaki Tsuchiya, Nobuhiko Tachibana, Kohjiro Nagao, Tomonori Tamura and Itaru Hamachi,“Organelle-selective click labeling coupled with flow cytometry allows pooled CRISPR screening of genes involved in phosphatidylcholine metabolism”,Cell Metabolism, Published online March 13, 2023, doi: 10.1016/j.cmet.2023.02.014
Contact
-
[About Research]
Itaru Hamachi
Professor, Graduate School of Engineering, Kyoto University
Tel:+81-75-383-2754
E-mail: ihamachisbchem.kyoto-u.ac.jp
Tomonori Tamura
Senior Lecturer, Graduate School of Engineering, Kyoto University
Tel:+81-75-383-2756
E-mail: tamurasbchem.kyoto-u.ac.jp
Masaki Tsuchiya
Assistant Professor, Graduate School of Engineering, Kyoto University
Tel:+81-75-383-2757
E-mail: tsuchiya.masaki.7akyoto-u.ac.jp
-
[About Program]
Go Kato
Department of Research Project, JST
E-mail: eratowwwjst.go.jp
