A Japanese research group developed a technology termed Photo-Isolation Chemistry (PIC) and successfully detected genes acting in very small cell populations and in intracellular microstructures by photoirradiation.
Humans and other multicellular organisms are known to be composed of at least 100 different types of cells, which are subdivided into smaller groups of cells with unique functions and characteristics depending on their spatial localizations. As these cells lose their intrinsic characters once they are isolated from an organ or tissue, analysis focusing exclusively on a specific type of cells has to be done without tissue destruction; however, conventional techniques did not have the capacity of meeting such a requirement.
A research group led by Professor Yasuyuki Ohkawa at the Medical Institute of Bioregulation, Kyushu University and Program-specific Associate Professor Shinya Oki at Graduate School of Medicine, Kyoto University developed a technology to isolate information on gene expression only from a photo-irradiated area, inspired by an ultrafine technology with photo-etching in a semiconductor manufacturing process. With this technology termed PIC, they photo-irradiated various areas in the brain and successfully detected the genes expressed in a specific domain of hippocampus. They also applied PIC to comprehensively detect the genes in very small cell populations of a mouse fetus, as well as those in intracellular microstructures smaller than one micron (1/1000 mm), which cannot be detected with conventional methods.
PIC is characterized as one of the technologies called “spatial transcriptomics”, where many researchers are competitively developing new technologies worldwide to link the gene expression with spatial information. Many of the spatial transcriptomics technologies require a large cost to use. In contrast, PIC requires only several dollars in addition to the cost for existing gene analysis techniques. In the future, PIC is expected to be used in pathological diagnosis involving clinical tissue specimens containing mixed normal and abnormal cells, such as diagnosis of cancer and inflammatory responses associated with COVID-19.
This research is conducted under the JST’s Strategic Basic Research Programs CREST (Team type) “Innovative Technology Platforms for Integrated Single Cell Analysis” and PRESTO (Individual researcher type) “Dynamics of Cellular Interactions in Multicellular System.”
-
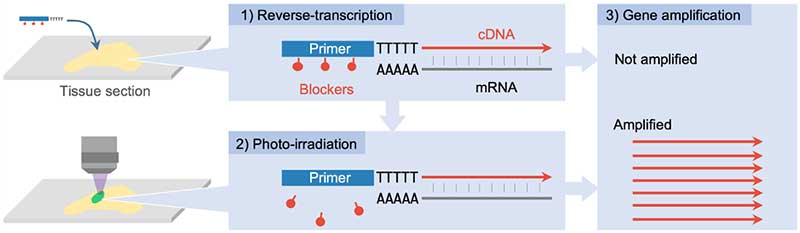
Figure 1. A principle of PIC technology to detect gene expression from photo-irradiated areas. 1) A primer with a photocleavable blocker is dropped onto the section. The RNA of the gene is then converted to DNA by the reverse transcription reaction. 2) The blocker is then removed from the primer when the target cells are photo-irradiated. 3) Only the gene from the photo-irradiated areas are amplified and detected. © Shinya Oki and Yasuyuki Ohkawa
-
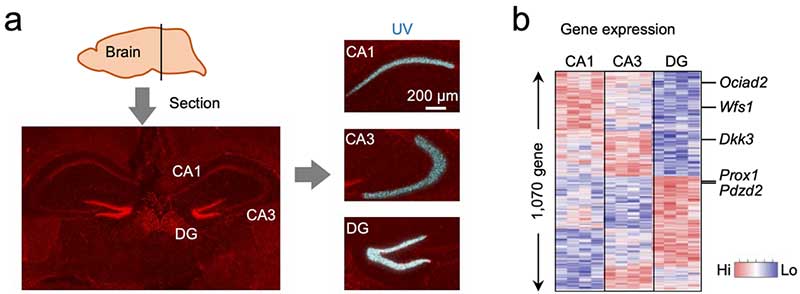
Figure 2. PIC for the hippocampus of mouse brains (a) Sections of the brain were prepared and PIC was performed by photo-irradiation to CA1, CA3, or dentate gyrus (DG) regions of the mouse hippocampus. (b) As a result, the expression levels of Ociad2 and Wfs1 in CA1, Dkk3 in CA3, and Prox1 and Pdzd2 in the DG were determined to be high, which is consistent with the results of a previous paper analyzing their gene expression patterns. © Shinya Oki and Yasuyuki Ohkawa
-
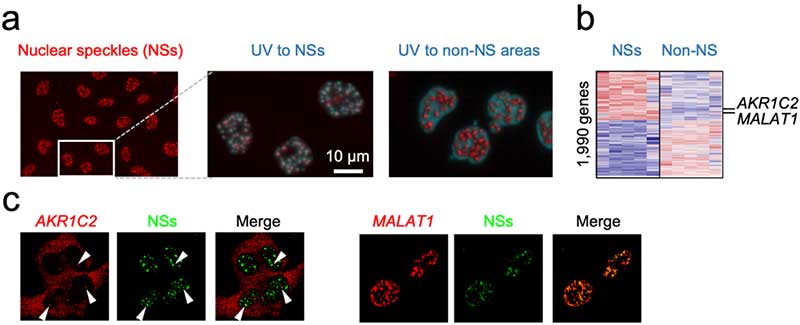
Figure 3. PIC for the nuclear speckles (a) PIC was performed by photo-irradiating to the nuclear speckles in the nucleus of the cell or to other nuclear regions. (b) As a result, AKR1C2 and MALAT1 genes were identified as abundant.
© Shinya Oki and Yasuyuki Ohkawa
Program Information
- JST CREST
- Research Area “Innovative Technology Platforms for Integrated Single Cell Analysis”
- Research Theme “The development of a high resolution technology in epigenomics”
- JST PRESTO
- Research Area “Dynamics of Cellular Interactions in Multicellular System”
- Research Theme “Elucidation of multicelluar system with spatial recording technology”
Journal Information
Mizuki Honda, Shinya Oki, Ryuichi Kimura, Akihito Harada, Kazumitsu Maehara, Kaori Tanaka, Chikara Meno and Yasuyuki Ohkawa, “High-depth spatial transcriptome analysis by photo-isolation chemistry” Nature Communications. Published online July 20, 2021,doi: 10.1038/s41467-021-24691-8
Contact
-
[About Research]
Yasuyuki Ohkawa
Professor, Medical Institute of Bioregulation, Kyushu University
TEL:+81-92-642-4531
E-mail: Yohkawabioreg.kyushu-u.ac.jp
Shinya Oki
Associate Professor, Graduate School of Medicine, Kyoto University
TEL:+81-75-366-7479
E-mail: oki.shinya.3wkyoto-u.ac.jp
-
[About Program]
Yasuda Mutsuko
Department of Strategic Basic Research, JST
E-mail: crestjst.go.jp
