Under JST’s Strategic Basic Research Programs, Noda Nobuo (Laboratory Head) and Yamasaki Akinori, Postdoctoral Fellow (currently Assistant Professor, Tokyo Institute of Technology), at the Institute of Microbial Chemistry in collaboration with other researchers, have discovered that autophagy is effective for selectively degrading protein in a state of liquid droplet1) that is formed through liquid-liquid phase separation2) but does poorly with the degradation of protein in aggregation or solid state.
Autophagy is one of the mechanisms through which cellular protein is degraded. There exists a selective type of autophagy named selective autophagy3) that targets and degrades specific proteins and organelles. It has been assumed that selective autophagy prevents the onset of diseases, but the state of proteins in which they could be efficiently degraded had been unclear.
This research group used the selective autophagy of Ape1 protein in yeast as the model system and succeeded in reconstituting the Ape1 isolation process with a lipid membrane in a test tube. It revealed that Ape1 protein is effectively isolated by the lipid membrane through the functioning of the Atg8 and receptor proteins when Ape1 forms droplets but is not isolated by the lipid membrane when Ape1 forms aggregates.
The discovery that selective autophagy is effective in degrading protein droplets but does poorly in degrading protein aggregates indicates that the activation of autophagy alone is insufficient in the prevention of and therapeutic medication development for neurodegenerative diseases and other diseases that are believed to be caused by the accumulation of abnormal proteins and that it is important to develop drugs that change aggregations into droplets. This will hopefully lead to the discovery of drugs that target “liquid-liquid phase separation,” which is the mechanism for creating droplets.
It was also proved for the first time that the isolation step for the targeted protein in selective autophagy can be conducted with just the Atg8 and receptor proteins. Autophagy also selectively degrades mitochondria and other organelles as well as pathogenic bacteria. Atg8 and receptor proteins could be responsible for selective isolation in a similar mechanism in these cases as well. The application of the test-tube reconstitution system developed here would promote research on the selective autophagy mechanisms for a wide variety of targets.
1) Liquid droplet
A condensate of macromolecules with fluidity created when protein and/or nucleic acids undergo liquid-liquid phase separation. The droplets are also known as “membraneless organelles” and perform various functions within the cell. A droplet spontaneously assumes a spherical form. It also has high internal fluidity, and it actively exchanges molecules with its surroundings.
2) Liquid-liquid phase separation
This is the phenomenon of a uniform liquid phase separating into multiple liquid phases. It is observed in daily life as the separation of water and oil, which occurs within cells with proteins and nucleic acids.
3) Selective autophagy
Autophagic pathway that degrades specific targets when it is required by the cell, unlike ordinary autophagy, which is triggered by starvation and degrades intracellular components indiscriminately. In selective autophagy, there are receptor proteins that recognize the wide variety of targets for degradation. It is believed that they all selectively load their targets into autophagosomes by connecting to both the degradation targets and the Atg8 proteins on the autophagosomes.
-
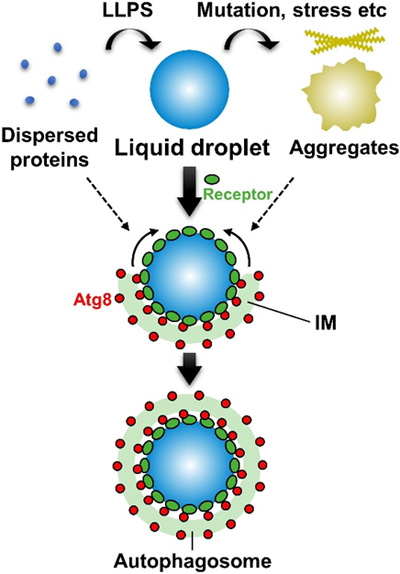
Figure. Selective autophagy efficiently degrades liquid droplets Some dispersed proteins undergo liquid-liquid phase separation (LLPS) to form liquid droplets. Liquid droplets possess high fluidity and are more similar to a liquid state than a solid state. It is known that liquid droplets lose their liquidity by mutation or stress and are transformed into aggregates and amyloids. Selective autophagy preferentially targets liquid droplets during which receptors accumulate on the surface of droplets and interacts with Atg8 on the isolation membrane (IM), thereby enabling the expansion of the IM along the surface of the droplets. On the other hand, selective autophagy is not good at degrading solid-like aggregates or dispersed proteins.
© Institute of Microbial Chemistry
Program Information
- JST CREST
- Research Area “Structural Life Science and Advanced Core Technologies for Innovative Life Science Research”
- Research Theme “Structural life science for unveiling membrane dynamics during autophagy”
Journal Information
Akinori Yamasaki, Jahangir Md. Alam, Daisuke Noshiro, Eri Hirata, Yuko Fujioka, Kuninori Suzuki, Yoshinori Ohsumi and Nobuo N. Noda. “Liquidity Is a Critical Determinant for Selective Autophagy of Protein Condensates”, Molecular Cell. Published online January 28, 2020, DOI: 10.1016/j.molcel.2019.12.026
Contact
-
[About Research]
Noda Nobuo
Laboratory of Structural Biology, Institute of Microbial Chemistry
TEL: +81-3-3441-4173
E-mail: nnbikaken.or.jp
[About Program]
Kawaguchi Tetsu
Department of Strategic Basic Research, JST
E-mail: crest
