The brain stores memories through a neuronal ensemble, termed engram cells. A unique system was established to transfer neuronal population activity into light with discrimination between engram and non-engram cells using fluorescence proteins. By using this system, it is revealed that engram sub-ensembles represent distinct pieces of information, which are then orchestrated to constitute an entire memory. In addition, some sub-ensembles preferentially reappear during post-learning sleep, and these replayed sub-ensembles are more likely to be reactivated during retrieval.
Japanese research group supervised by Dr Noriaki Ohkawa (Lecturer) and Professor Kaoru Inokuchi of University of Toyama succeeded to establish a system to investigate characteristic activity of cell ensemble acquiring memory and visualized ways for representation and consolidation of memory experienced novel episode in brain.
We are exposed to many episodic events and then memorize their information through our life. This kind of memory, episodic memory, is processed in several brain regions, and one of the regions is hippocampus. It is authorized that, in the hippocampus, one specific episodic memory is stored within and retrieved from a neuronal ensemble composed of neurons, termed engram cells, that are activated during learning. Indeed, activation or inhibition of engram cell ensemble induces or inhibits memory retrieval, respectively, thus, engram cell ensemble represents the physiological manifestation of a specific memory trace. However, one episodic memory is composed of several components of episode, and each component should be encoded by specific substrate, e.g. engram sub-ensemble. Nevertheless, it had not been clear how activity in these engram cells is assembled to represent the corresponding event because of technical limitations mean that it is difficult to distinguish the population activity of engram cells from that of non-engram cells.
To address how one episodic memory is represented and consolidated in engram cell ensemble, it is required to visualize the activity of engram and non-engram cells. Engram cells can be specifically targeted in c-fos-tTA mice because the neural activity associated with memory formation induces c-fos gene expression, which in turn induces activity-dependent tTA expression under the control of the c-fos promoter. In the absence of doxycycline, tTA can bind to the tetracycline responsive element (TRE), enabling downstream expression of the TRE-dependent transgene (Figure 1a). When neuron activates, Ca2+ flows into their soma. Thy1-G-CaMP7 mice express a Ca2+ indicator, G-CaMP7, in pyramidal neurons of hippocampal CA1 in the mice. Thus, neuronal activity is transferred into G-CaMP7 fluorescence, called Ca2+ imaging. We developed a technique that combines a head-mounted, miniature fluorescent microscope, with Thy1-G-CaMP7/c-fos-tTA double transgenic mice. The hippocampal CA1 region in double transgenic mice was injected with LV expressing a fluorescent protein, Kikume Green Red (KikGR), under the control of TRE (Figure 1a and 1b). Using this approach, engram cells can be identified with KikGR, and the Ca2+ signals corresponding to the activity of engram cells and non-engram cells can be tracked during experience of a novel episodic event (Figure 1c and 1d).
The data indicated that population activity of engram cell ensemble exhibited the characteristic trait of highly repetitive activity during novel episodic event. To address component of one memory, next, it was proposed to deconstruct population activity into sub-ensemble groups. Non-negative matrix factorization (NMF) decomposes population activity into a time series of coactivated neuronal ensembles (Fig. 2a). Each sub-ensemble is composed of the different cells, which are spatially intermingled (Fig. 2b left), to make their synchronous activity even among the group of engram cells associated with a single event (Fig. 2b right). These results suggest that the total information of one event is structured into sub-engram ensembles.
In order to measure the activity of engram cells across different memory processing stages, recording of Ca2+ transients from novel experience through post-experience sleep to retrieval was conducted. Around 40% of the total number of sub-ensembles that appeared in a novel experience in engram reactivated during post-experience sleep and then preferentially reappeared in retrieval sessions, while almost none of sub-ensembles in non-engram did not show this feature. Thus, engram sub-ensembles formed during a novel experience and that were reactivated during sleep sessions were mostly reactivated during the retrieval session (Fig. 3). By contrast, most non-engram ensembles that were activated during the novel experience were not reactivated in the later sessions.
The findings reported in this study demonstrate that engram cells possess synchronous activity, formed by several sub-ensembles in the engram population. Only in engram cells does this synchronous activity survive through post-learning sleep sessions that contribute to the consolidation process. The present work sheds light on the relationship between ensemble activities and coding principles in learning and memory.
-
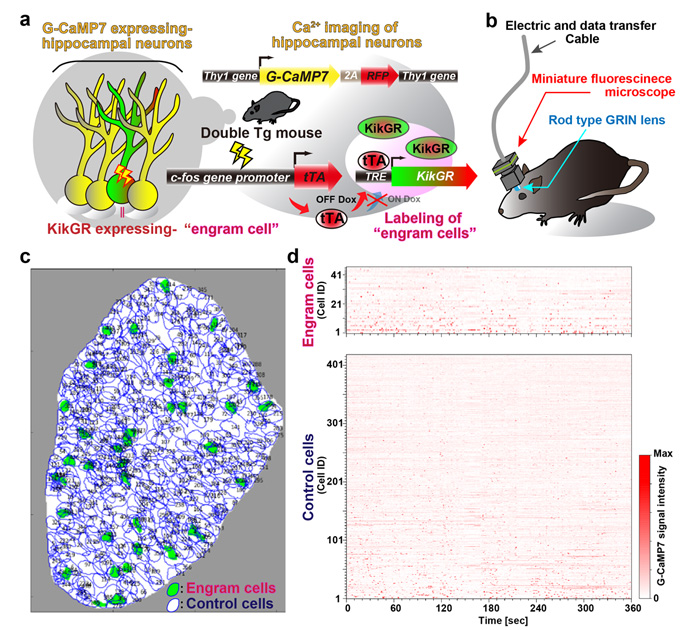
Figure 1. New optogenetic system to reveal neuronal activity of engram and non-engram ensemble by light. - a. Engram cells can be specifically targeted in c-fos-tTA mice. Thy1-G-CaMP7 is useful for Ca2+ imaging of hippocampal CA1 neurons. Sander symbol, indicating neuronal activation state.
- b. Image of observation of fluorescence in mouse brain with miniature fluorescent microscope.
- c. Representative image of all engram cells and non-engram control cells detected from a single animal.
- d. Normalized calcium transients corresponding to neuronal activity for all engram cells (top) and non-engram cells (bottom) during experience of novel episode (displayed over time).
-
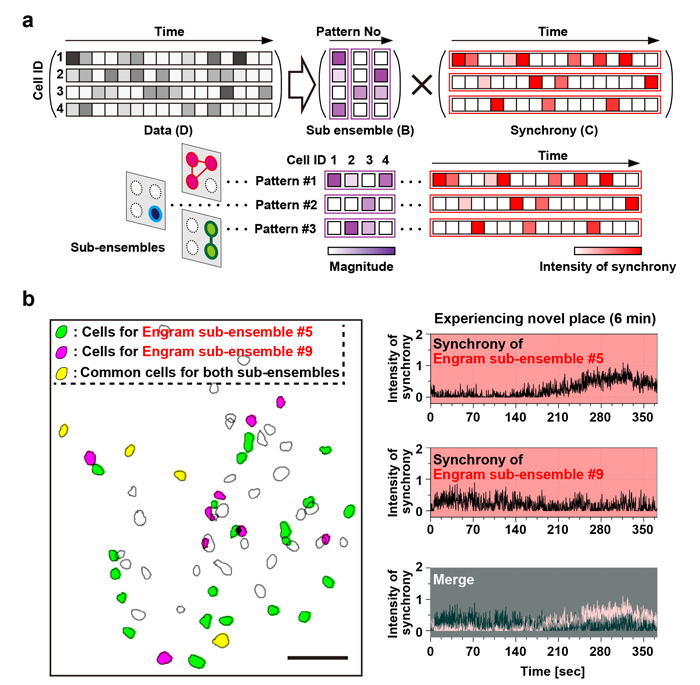
Figure 2. Episodic memory of experience of novel space is constructed by several sub-ensembles within a single engram population. - a. Diagrammatic representation of NMF analysis; colored bars indicate the relative activity of each neuron in the detected pattern (shown as the magnitude) (left), and the relative intensity of the synchronous activity of this pattern across time (right).
- b. Example of two detected engram sub-ensembles. Distribution map of detected engram cells (left panel). Green and red magenta filled contours indicate neurons assigned to two different ensembles (corresponding to patterns #5 and #9, respectively), and yellow filled contours denote cells shared between patterns. Scale bar, 100 µm. The temporal pattern of synchronous activities of patterns #5 and #9 during experiencing novel space (top and middle right) and their overlay (bottom right).
-
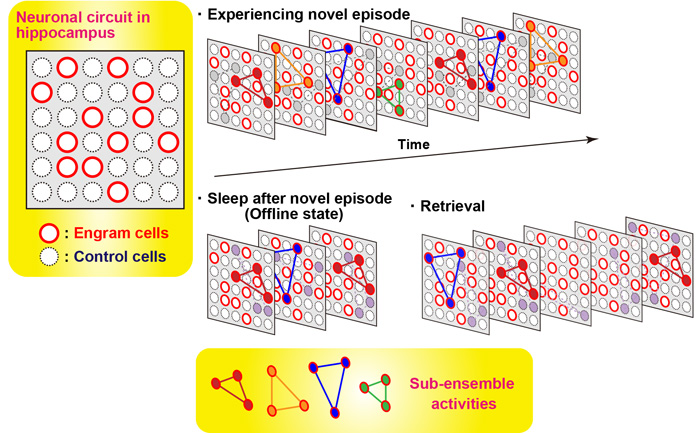
Figure 3. Diagrammatic model showing orchestration of ensembles of engram cells across memory processing. Many sub-ensembles express during experiencing novel episode. A part of sub-ensembles of engram cells re-appear during post-episode experience sleep. This set of sub-ensembles appeared during sleep are more likely observed during memory retrieval. In contrast, control, non-engram, cells almost do not show this feature.
Program Information
- JST CREST
- Research Area “Creation of Fundamental Technologies for Understanding and Control of Biosystem Dynamics”
- Research Theme “Memory integration process based on cell assembly and neural circuit model”
- JST PRESTO
- Research Area “Development of optical control technologies and elucidation of biological mechanisms”
- Research Theme “Creation of new methods for manipulation of memory by extraction of characteristic patterns of engram”
Journal Information
Khaled Ghandour, Noriaki Ohkawa, Chi Chung Alan Fung, Hirotaka Asai, Yoshito Saitoh, Takashi Takekawa, Reiko Okubo-Suzuki, Shingo Soya, Hirofumi Nishizono, Mina Matsuo, Makoto Osanai, Masaaki Sato, Masamichi Ohkura, Junichi Nakai, Yasunori Hayashi, Takeshi Sakurai, Takashi Kitamura, Tomoki Fukai & Kaoru Inokuchi. “Orchestrated ensemble activities constitute a hippocampal memory engram”, Nature Communications. Published online June 14, 2019, DOI: 10.1038/s41467-019-10683-2
Contact
-
[About Research]
Kaoru InokuchiProfessor, Department of Biochemistry, Graduate School of Medicine and Pharmaceutical Sciences, University of Toyama
E-mail: inokuchimed.u-toyama.ac.jp
-
[About Program]
Tetsu Kawaguchi
Department of Strategic Basic Research, JST
E-mail: prestojst.go.jp
